Grouping sequencing into motu using abgd
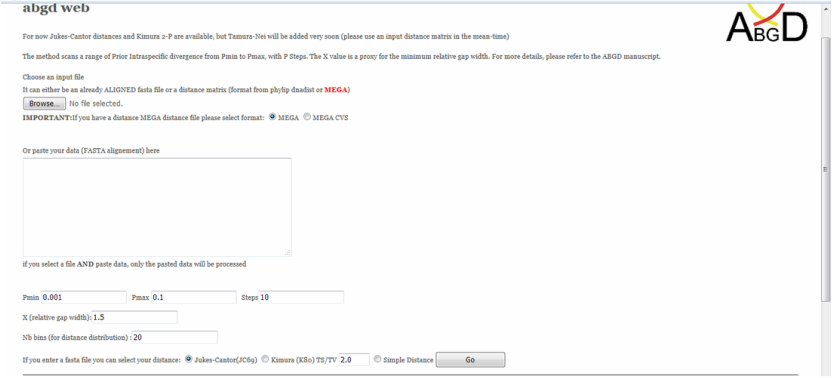
To group sequences into MOTU (molecular operational taxonomic units), a popular tool is the ABGD (automatic barcode gap discover). The online ABGD is available here: http://wwwabi.snv.jussieu.fr/public/abgd/
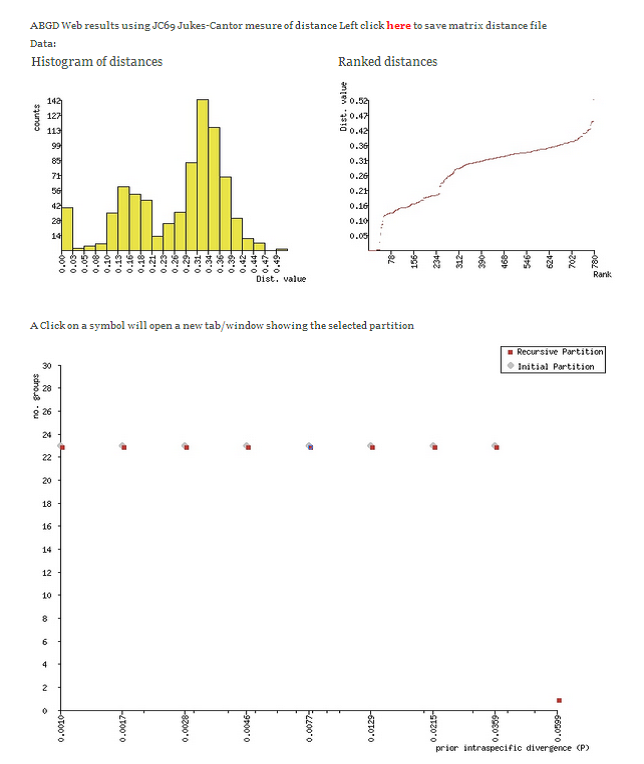
ABGD uses an automatic recursive procedure to converge on the best patterns for the dataset and arranges DNA barcodes into clusters (partitions) accordingly. The median number of ABDG partitions can be used as the basis for MOTU (species) as this has produced good correspondence with traditional species in empirical studies.
BOLD also has its own MOTU grouping algorithm, and DNA barcodes uploaded to BOLD (provided they meet a certain threshold of length and quality will be sorted into BINS (Barcode Index Numbers).